SNP Polymorphysim Microarray Chip - How to Test a Person's DNA
By: HWC
Date Uploaded: 06/03/2020
Tags: homeworkclinic.com Homework Clinic HWC test a person's DNA blood sample lack nuclei white blood cells red blood cells chromosomal DNA homozygous heterozygous nucleotide polymorphism SNP polymerase chain reaction DNA chip fluorescent color polymorphism SNP Polymorphysim Microarray Chip animation
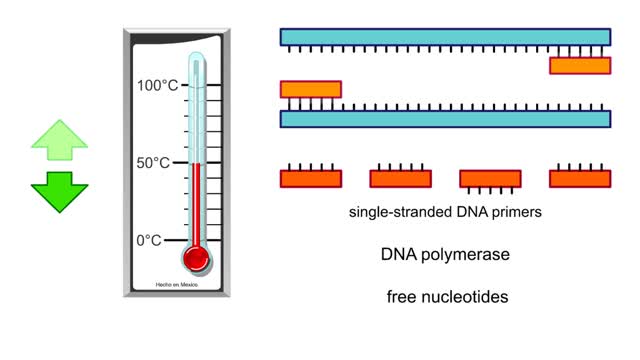
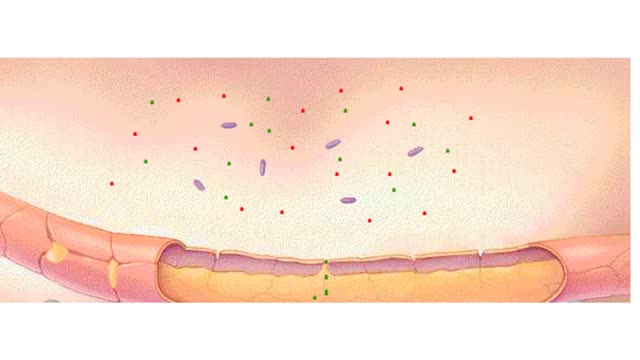
To test a person's DNA, a researcher first needs a source of tissue. Most of the cells in a blood sample are red blood cells, which lack nuclei, but there are also a number of white blood cells, which do contain nuclei and chromosomal DNA. If we could see a particular DNA sequence in these cells, we would identify two copies of the sequence, because the cells are diploid and have inherited one DNA copy from each parent. In the example above, individuals 1 and 3 are homozygous for a particular sequence, while individual 2 is heterozygous for this sequence. Let's say that the sequences differ by a single nucleotide. In one sequence, a C exists in a particular position, and in the other sequence, the C is replaced by a T. This difference is a single nucleotide polymorphism, or SNP. Before a researcher can identify the SNPs, the DNA must be copied so that the researcher has enough DNA to work with. The sequences are copied using the polymerase chain reaction, or PCR. The researcher can then either sequence the DNA directly or subject the sample to DNA chip analysis. Here we examine how researchers use DNA chips to identify SNPs. The DNA strands produced by PCR are separated and applied to the DNA chip. These chips are designed so that they have short pieces of DNA permanently affixed to them. The sequences of the short pieces of DNA are complementary to the DNA containing the SNPS. Notice that the PCR products from the heterozygous individual differ at one position, either containing a C or a T. The short DNA sequences end just before the SNP in question. If a DNA polymerase enzyme is added to the chip, it will incorporate a nucleotide that is complementary to the SNP into the end of the short strand. For this type of experiment, the nucleotides to be incorporated are first labeled with fluorescent color tags. In this heterozygous sample, a G has been incorporated into one strand, while an A has been incorporated into the other. Whereas the heterozygous sample contains two colors, the two homozygous samples each contain just one color. Using this technique, researchers can tell what SNPs a person has, and whether the person is homozygous or heterozygous. In this heterozygous sample, a G has been incorporated into one strand, while an A has been incorporated into the other. Whereas the heterozygous sample contains two colors, the two homozygous samples each contain just one color. Using this technique, researchers can tell what SNPs a person has, and whether the person is homozygous or heterozygous. In the example of SNP2, in which a different sequence of DNA is analyzed, individual 1 is homozygous for a particular polymorphism. This polymorphism is represented by green. Individuals 2 and 3 are homozygous for a different polymorphism, represented by blue.
Add To
You must login to add videos to your playlists.
Advertisement






Comments
0 Comments total
Sign In to post comments.
No comments have been posted for this video yet.